Introduction
Diversity is very important. Understanding why diversity is lost is important to understanding the overall health of ecosystems. Observe the major difference in diversity between a grassland and a forest habitat; the forest habitat is much more diverse. Forests are not homogeneous throughout, i.e. diversity increases as you move from the edge to the middle of the forest. Forest edges receive more attention thanks to human activity forming more edges. This habitat fragmentation can reduce diversity among species (Beasley, 2019).
Using a systematic sampling transect we can understand the changes in diversity from the forest edge to the middle of the forest. A transect is a process that takes samples along a straight line with a goal to observe the changes in diversity along that line. The transect line can vary in length, anywhere from a few meters long to a hundred meters long. There are two kinds of transect sampling methods: a line transect where the individuals touching a meter length of string stretched along the transect are recorded; and a belt transect where quadrants are placed at intervals along the transect and organisms in each quadrant are recorded. The line transect is more convenient but generates less complete data while the belt transect is more strenuous but generates more complete data (Beasley, 2019).
Method
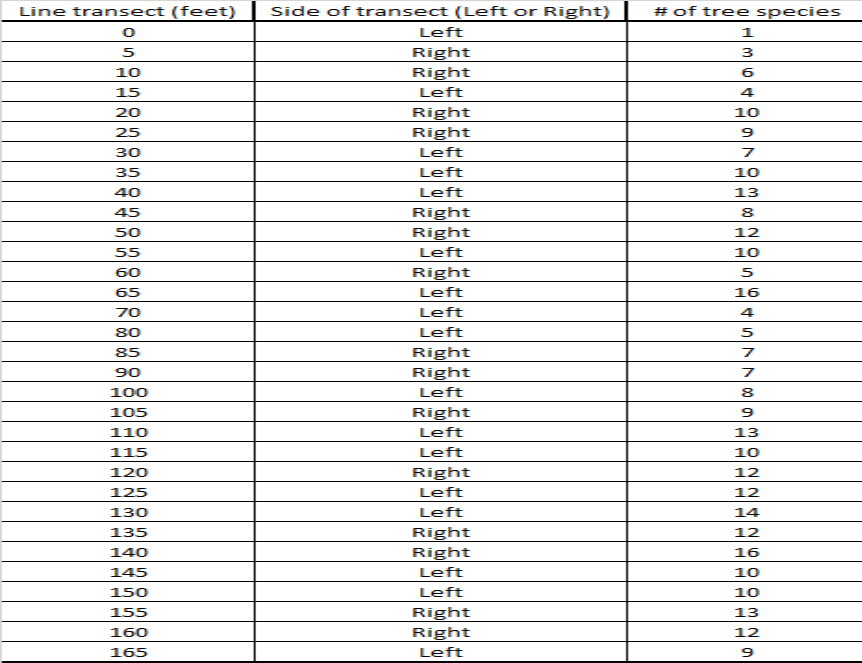
For our lab we used the line transect sampling method to assess tree species diversity from the forest edge to the middle of the forest at Stringers Ridge, an urban forest, here in Chattanooga, TN. Our group members were Will Foushee, Luke Mainello, and myself. We used a transect tape to make a 50 meter (165 ft) line from the hiking trail to the forest interior. We then tracked the amount of different tree species every 1.5 meters (5 ft) using the string provided. We determined which side of the transect line to sample by flipping a coin. We were able to identify the tree species by using the tree guide provided. Afterwards we used the data to make a scatter plot showing change in tree species diversity starting at zero m, the forest edge, to 50 m, forest interior.

Results
Analysis of the data collected show a small, non-significant relationship between the forest gradient (distance from forest edge to forest interior) and species richness. Our R^2 value of 0.31 is closer to 0 than it is to 1.

Discussion
1. Provide an overall description of the habitat. Some characteristics you may consider include: soil condition, evidence of disturbance, temperature, light intensity, tree size, water condition, etc. The habitat varied in elevation, i.e. it was very sloped. The soil appeared to be dry and compact. There was evidence of disturbance, e.g. fallen trees, but there’s no way to discern what caused the disturbance. The disturbance did not appear to be human caused. The temperature was high enough to make me roll my sweater sleeves up, so mid- to high 70’s (degrees Fahrenheit). Light intensity was strong at the beginning of the trail, but where we started our transect it was fair. The end of our transect had minimal light intensity. Trees varied in size; most were samplings and the largest had a diameter of only 2 to 3 ft. The habitat appeared to have minimal water signs; other than the growth of the trees.
2. Based on your scatter-plot, what pattern did you observe in tree species diversity as you moved from the edge of the forest to the interior? Does the pattern have a strong or weak effect on number of tree species? How do you know? The scatter-plot showed a non-significant relationship between the forest gradient and the species richness, but it obviously shows that diversity rose the further we went into the forest interior. The weak pattern is shown by the r^2 value of 0.31, which is closer to 0 (which shows no pattern) than 1 (which shows a definite pattern).
3. Run a regression analysis in Microsoft Excel on your data. In Data Analysis, select regression. Select the line transect values for the Input X range and number of tree species for the Input Y range. Explain whether or not the relationship is statistically significant. After running the regression analysis, a resulting p-value of 3.63E-06 was displayed. Because the p-value is so low the relationship is not statistically significant.
4. As the terrestrial environment becomes increasingly fragmented, what patterns in tree diversity might you expect to see based on your results? What follow up questions do you have for a future study? Based on my results, I believe that diversity patterns in trees will slowly decline with more habitat fragmentation. To follow up I would ask, “What recent events, biotic and abiotic, could have disrupted the community?”
Read the article, “Gene Flow Halted by Fragmented Forests”
1. According to the article, why is the conservation of river floodplain ecosystems important? Because river floodplain ecosystems help maintain water quality. They also help prevent erosion and provide important habitat for wildlife (Saeki, 2018).
2. Explain why gene flow is important for monitoring endangered species such as Acer miyabei. How does landscape genetics help to understand gene flow patterns? Because less gene flow between populations will possibly result in the disruptive (i.e. diversifying) natural selection which favors two or more extreme phenotypes over the average phenotype. This disruptive selection could lead to two or more distinct sub-species. Landscape genetics helps to understand gene flow patterns by analyzing the DNA of similarly aged species and comparing the different ages’ DNA (Saeki, 2018).
3. What were the overall findings of the study? How does such research inform our conservation and restoration efforts? The overall finding was that the older trees had more similar phenotypes because they once coexisted in the same habitat, and that younger trees’ DNA were more varied do to isolating populations from habitat fragmentation. This research helps inform scientist the overall effects of habitat fragmentation on species’ DNA and the changes the DNA are evolving towards (Saeki, 2018).
Cited References
Saeki, I., Hirao, A. S., Kenta, T., Nagamitsu, T., & Hiura, T. (2018). Landscape genetics of a threatened maple, Acer miyabei: Implications for restoring riparian forest connectivity. Biological Conservation,220, 299-307. doi:10.1016/j.biocon.2018.01.018